 
Note that the normalized count values are log2.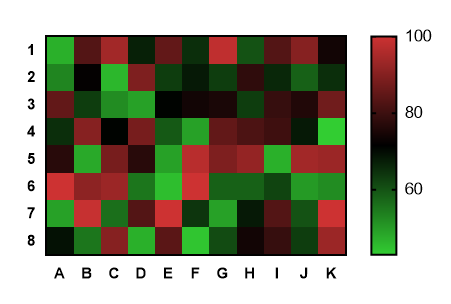 
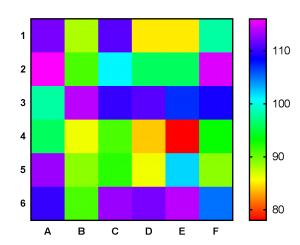
It should look like below (just the first few rows and columns are shown). Next we need to get the normalized counts for these genes, from the file containing the normalized counts for all genes in the experiment, and then extract just the columns we need for the heatmap (the normalized counts and gene labels).įirst click on the galaxy-eye (eye) icon and take a look at the normalized counts file that we imported. Now we have a file that contains only the top 20 genes from the DE results. param-text “Select first”: 21 (20 genes plus header row)Įxtract the normalized counts for top genes. 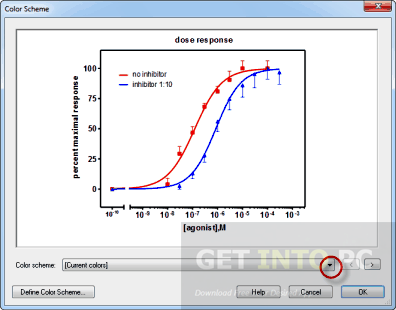
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
